> x1 <- c(3,2,5,1,4) > x2 <- c(6,4,5,7,2) > t.test(x2, mu=10, alternative="less") # 1-way 1 sample t-test > t.test(x1, x2) # 2 sample t-test > help(t.test) > wilcox.test(x1,x2, paired=T) # matched-pairs Wilcoxon test
> plot(iris$Petal.Width, iris$Petal.Length) > fit.i1 <- lm(Petal.Length ~ Petal.Width, data=iris) > coef(fit.i1) > summary(fit.i1) > abline(coef(fit.i1)) > pred1 <- predict(fit.i1, interval="confidence") # narrow confidence band > pred2 <- predict(fit.i1, interval="prediction") # wide prediction band > petW <- iris$Petal.Width > matlines(petW, pred1, col="red") > matlines(petW, pred2, col="blue") > plot(fitted(fit.i1), resid(fit.i1))
> av <- lm(Petal.Length ~ Species, data=iris) # 1-way Anova > anova(av) > summary(av) > boxplot(Petal.Length ~ Species, data=iris) > with(iris, stripchart(Petal.Length ~ Species, "jitter", vert=T))Make sure the independent variable is a factor. If it is numerical (e.g. 1, 2, 3), convert the column to factors:
> dat$x <- factor(dat$x)Without the conversion, you are doing linear regression.
The traditional 1-way ANOVA assumes the equal variance for all treatment groups. The following test (Welch 1951) relaxes this assumption:
oneway.test(Petal.Length ~ Species, data=iris)
- Notation:
- y, x1, and x2 are numeric variables.
- A, B, and C are factors.
y ~ xory ~ x + 1
Simple linear regression of y on x with intercept: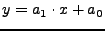 .
.
y ~ x - 1
Simple linear regression without intercept; i.e., force the line to go through the origin: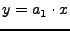
log(y) ~ x1 + x2
Multiple regression with log-transformed y. Implicitly, the intercept is included.y ~ x + I(x^2)
Polynomial regresion: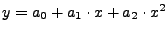
y ~ A + B
Two-way ANOVA without interaction termy ~ A + B + A:Bory ~ A*B
Two-way ANOVA with interaction termy ~ A*B*Cory ~ A + B + C + A:B + B:C + A:C + A:B:C
Three-way ANOVA with all possible interactionsy ~ A*B*C - A:B:C
Three-way ANOVA without the 3-way interaction (A:B:C) term.y ~ A + x
Single classification ANCOVA with class determined by A and with covariate xy ~ A * x
ANCOVA with class determined by A. Slopes are obtained for each class. x
ANCOVA example
> fit.cov1 <- lm(Petal.Length ~ Sepal.Length + Species, data=iris) > anova(fit.cov1) > fit.cov2 <- lm(Petal.Length ~ Sepal.Length * Species, data=iris) > anova(fit.cov2) > summary(fit.cov2)
- glm
- Generalized linear model, includes logistic regression
- lme
- Linear Mixed-effects Models. It is a part of package nlme ``non-linear mixed effects models''. You need to load the package by:
> library(nlme) > help(lme)
- Surv
- Survival analysis. Need to load survival package