A little bit of explanation and summary:
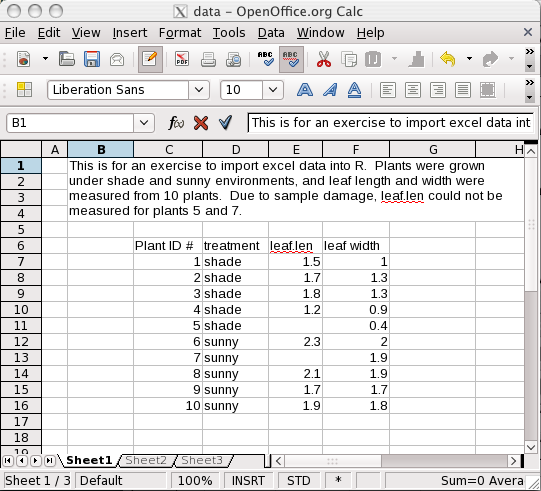
What we have done above is that you used the command (function)
read.table(), and the results of this command (data read in
the way R can understand) is stored in a container called
dat.in.
<- is called an assignment (it looks like an arrow). The results
of read.table() is assigned to dat.in.
You can use whatever name for this storage container (I chose
dat.in). Also, R can import multiple files. For example if
you have two files to read in, you can store one set of data in
simDat and the othe set in obsDat.
This storage container of spread-sheet like data is called
data frame, which we will talk more in the next section.