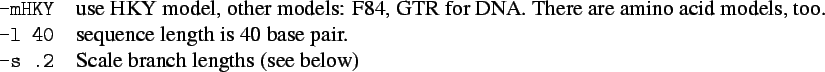
Traditional way of modeling the stochastic process is chronological order (forward modeling, e.g., diffusion model).
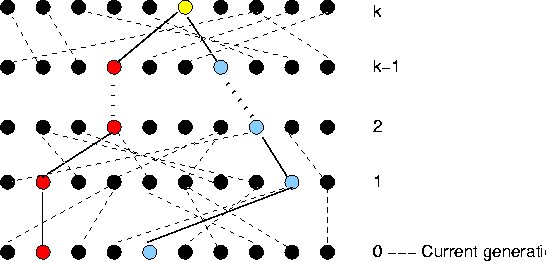
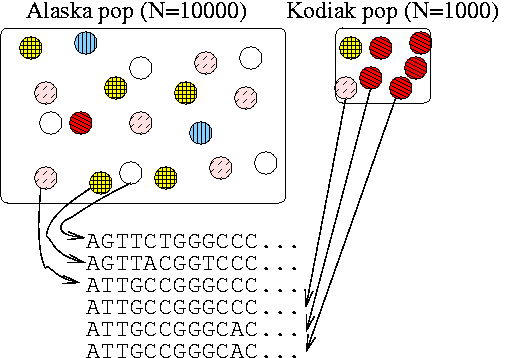
- Randomly pick 2 alleles from a population
- Go back in time until you find the most recent common ancestor (MRCA).
- Repeat this many times, and then take the average time to the MRCA.
- It is easy to calculate this expected time to the MRCA:
expected time to the MRCA of 2 alleles
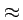 2 N
2 N
- If you compare two randomly sampled DNA sequences, how many
differences (
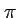 ) are there on average?
) are there on average?
- Expected time of coalescence among m samples
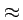 4
N
4
N
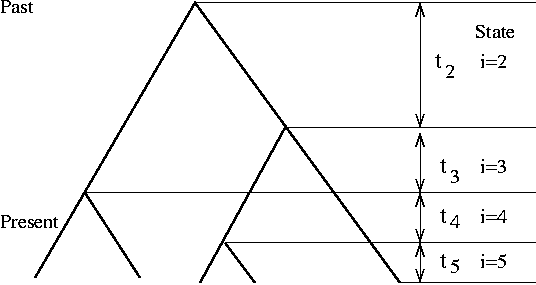
- Expected total branch lengths:
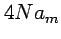 , where
, where 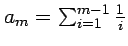 (Watterson's coefficient).
(Watterson's coefficient).
- Number of segregating sites:
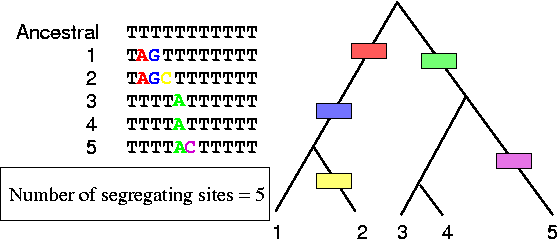
Expected number of segregating sites:
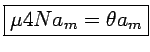
- Tajima's D
- Tajima's test statistics
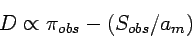 Note that the two elements are the estimaters of
Note that the two elements are the estimaters of 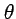 :
: 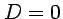 expected under the neutral evolution.
expected under the neutral evolution.
- It is a summary statistics of genealogical shape.
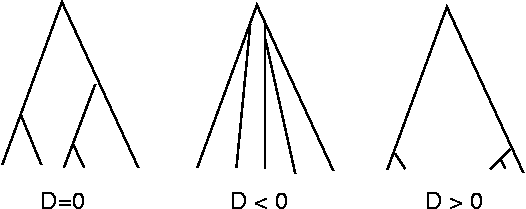
- Tajima's test statistics