In simulations, you frequently need to generate random number following a distributions other than uniform distributions. In previous exercises, you have already created a rather simple way to draw a random number from the binominal distribution and geometric distribution (remember the coin flipping functions in the homeworks?).
Here I briefly discuss about approaches to create non-uniform random numbers from uniform PRNG.
In reality, I generally use pre-made functions from a library called GNU Scientific Library (GSL).
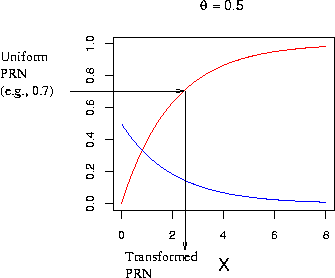
e.g., To create a random number y from exponential
distribution, draw a unifrom PRN z [0,1), and use the following
transformation.
#include <stdio.h>
#include <gsl/gsl_rng.h>
#include <gsl/gsl_randist.h>
const gsl_rng *gBaseRand; /* global rand number generator */
int main (void) {
unsigned long randSeed;
int i, k, n = 10;
double lambda = 3.0;
/* specifying to use Mersenne twister MT-19937 as the uniform PRNG */
gBaseRand = gsl_rng_alloc(gsl_rng_mt19937);
srand(time(NULL)); /* initialization for rand() */
randSeed = rand(); /* returns a non-negative integer */
gsl_rng_set (gBaseRand, randSeed); /* seed the PRNG */
/* print n random variates chosen from the poisson distribution with mean
parameter lambda */
for (i = 0; i < n; i++) {
k = gsl_ran_poisson (gBaseRand, lambda);
printf (" %u", k);
}
printf ("\n");
gsl_rng_free(gBaseRand);
return 0;
}
const gsl_rng *gBaseRand;
Making a special global variable (I'm calling it gBaseRand), which stores what kind of uniform random number generator you want to use.
gBaseRand = gsl_rng_alloc(gsl_rng_mt19937);
We are setting this variable to use gsl_rng_mt19937.
There
are many other PRNGs which you can specify (e.g., you can use the
familiar drand48() if you specify gsl_rnd_rand48).
gsl_rng_set (gBaseRand, randSeed);
Seeding the specified PRNG (gBaseRand) with a seed (randSeed).
k = gsl_ran_poisson (gBaseRand, lambda);
Use gBaseRand as the base uniform PRNG, and draw a random number from a poisson distribution (whose parameter is specified as lambda).
gsl_rng_free(gBaseRand);
Free the memory allocated for the PRNG
gcc prng.c -lgsl -lgslcblas -lm
The first argument for all the functions are the variable containing the type of uniform PRNG (here I'm using gBaseRand).
gsl_ran_binomial (gBaseRand, double p, unsigned int n)
gsl_ran_geometric (gBaseRand, double p)
gsl_ran_negative_binomial (gBaseRand, double p, double n)
gsl_ran_poisson (gBaseRand, double lambda)
Returns unsigned long int
gsl_rng_uniform_int(gBaseRand, unisigned long int n)
Uniform
distribution, between 0 to n-1 inclusive.
gsl_ran_gaussian (gBaseRand, double sigma) |
Normal distribution with mean of 0 |
gsl_ran_exponential (gBaseRand, double theta) |
Exponential distribution |
gsl_ran_flat (gBaseRand, double a, double b) |
Uniform (flat) distribution, a <= x < b, [a, b) |
gsl_rng_uniform(gBaseRand) |
Uniform, 0 <= x < 1, [0, 1) |
gsl_rng_uniform_pos(gBaseRand) |
Uniform, 0 < x < 1, (0, 1) |
There are many more distributions in GSL, and you can use them in the same way as the example above.
$ R
> dat <- read.table("out1.txt")
> hist(dat[,1])
> mean(dat[,1])
> var(dat[,1])
> q()
Compare the observed mean and variance with the expectation if you are using one of the distribution we covered in the class.
Try to use several different distributions. Also change the parameters of distributions, and observe how the shape changes.